Section:
New Results
Multi-scale Mining of fMRI Data with Hierarchical Structured Sparsity
Inverse inference, or "brain reading", is a recent paradigm for
analyzing functional magnetic resonance imaging (fMRI) data, based on
pattern recognition tools. By predicting some cognitive variables
related to brain activation maps, this approach aims at decoding
brain activity. Inverse inference takes into account the multivariate
information between voxels and is currently the only way to assess
how precisely some cognitive information is encoded by the activity
of neural populations within the whole brain. However, it relies on a
prediction function that is plagued by the curse of dimensionality,
as we have far more features than samples, i.e., more voxels than
fMRI volumes. To address this problem, different methods have been
proposed. Among them are univariate feature selection, feature
agglomeration and regularization techniques. In this work, we
consider a hierarchical structured regularization. Specifically, the
penalization we use is constructed from a tree that is obtained by
spatially constrained agglomerative clustering. This approach encodes
the spatial prior information in the regularization process, which
makes the overall prediction procedure more robust to inter-subject
variability. We test our algorithm on a real data acquired for
studying the mental representation of objects, and we show that the
proposed algorithm yields better prediction accuracy than reference
methods.
See also [29] and Fig. 6 .
Figure
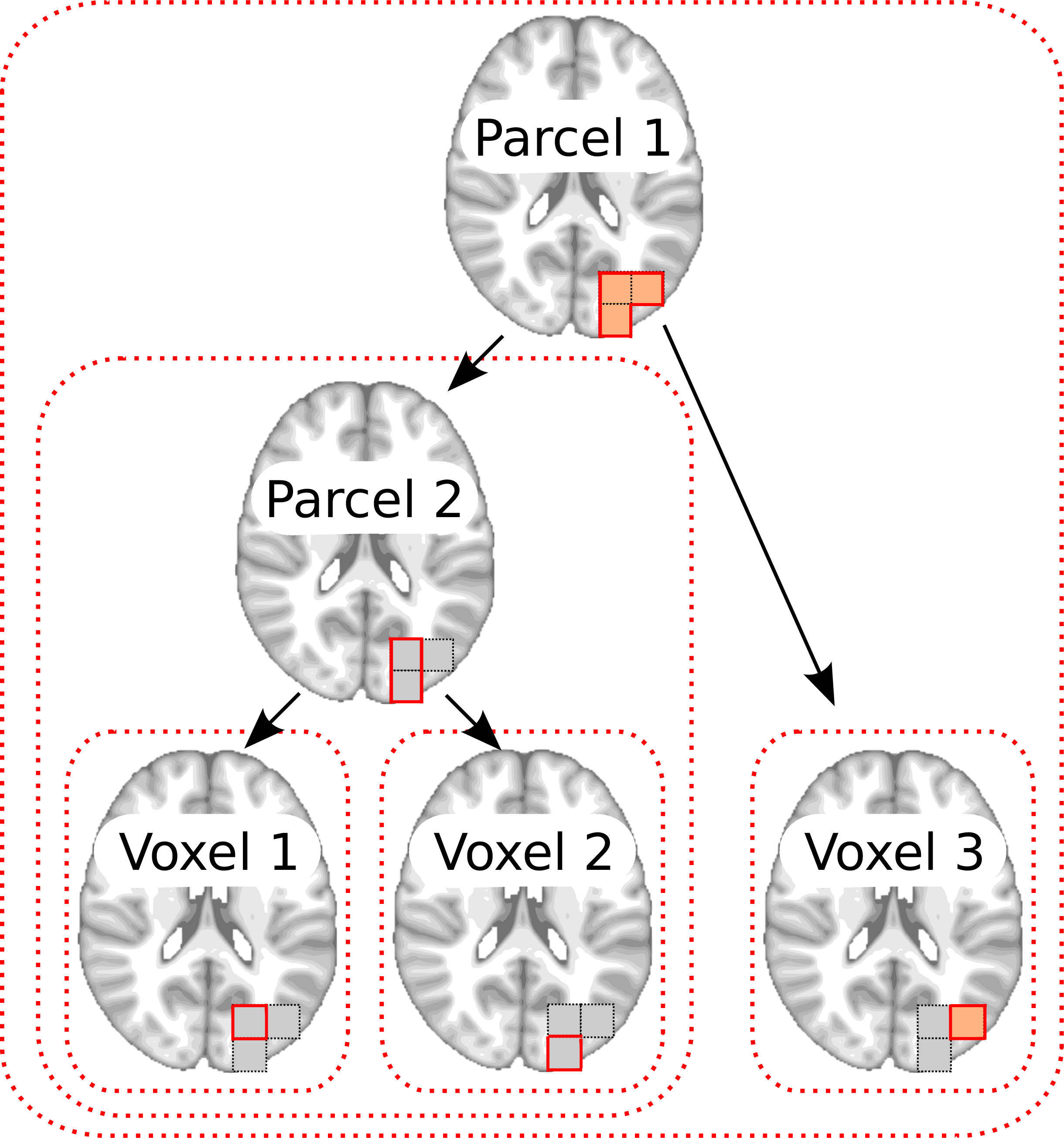
6. Principle of structured sparsity: Example of a tree
when , with three voxels and two parcels.
The parcel 2 is defined as the averaged intensity of the voxels
, while the parcel 1 is obtained by averaging the
parcel 2 and voxel 3. In red dashed lines are represented the
five groups of variables that compose . If the
group containing the parcel 2 is set to zero, the voxels
are also (and necessarily) zeroed out. |